Challenges of Intracellular Visualization using Virtual and Augmented Reality
Description:
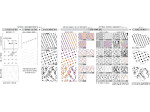
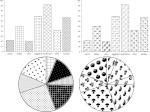
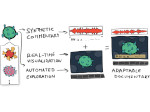
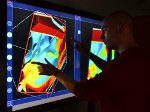
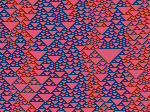
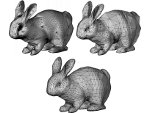
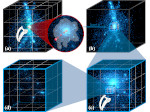
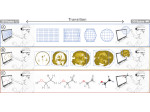
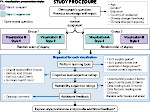
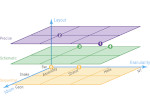
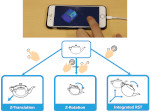
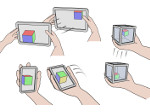
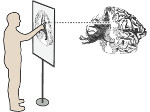
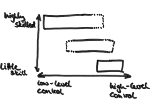
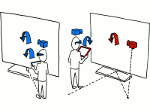
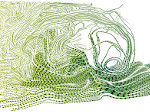
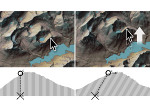
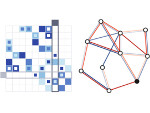
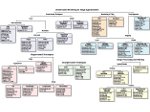
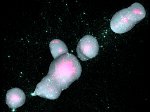
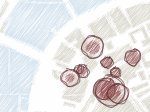
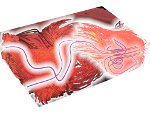
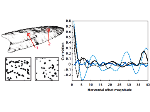
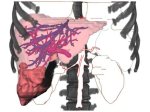
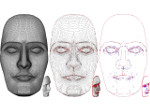
Microscopy image observation is commonly performed on 2D screens, which limits human capacities to grasp volumetric, complex, and discrete biological dynamics. With the massive production of multidimensional images (3D + time, multi-channels) and derived images (e.g., restored images, segmentation maps, and object tracks), scientists need appropriate visualization and navigation methods to better apprehend the amount of information in their content. New modes of visualization have emerged, including virtual reality (VR)/augmented reality (AR) approaches which should allow more accurate analysis and exploration of large time series of volumetric images, such as those produced by the latest 3D + time fluorescence microscopy. They include integrated algorithms that allow researchers to interactively explore complex spatiotemporal objects at the scale of single cells or multicellular systems, almost in a real time manner. In practice, however, immersion of the user within 3D + time microscopy data represents both a paradigm shift in human-image interaction and an acculturation challenge, for the concerned community. To promote a broader adoption of these approaches by biologists, further dialogue is needed between the bioimaging community and the VR&AR developers.
Paper download: 
 (1.8 MB)
(1.8 MB)
Software (from the sub-project that I was heading):
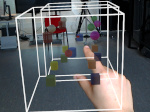
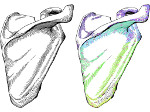
The AR software for exploring the HoloTracks visualization of live RPE1 cell stacks can be found in InraE's GitLab server: https://forgemia.inra.fr/nathan.van-hille/mrtk.
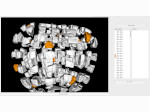
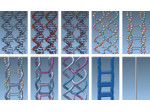
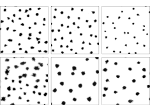
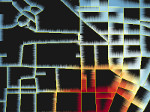
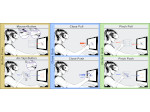
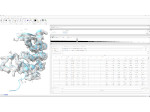
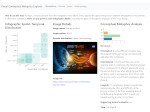
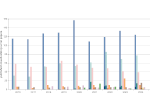
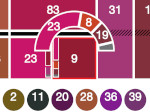
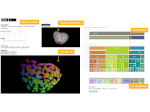
Pictures (from the sub-project that I was heading):
(these images as well as others from the paper, just like the whole paper, can be used under a  CC-BY 4.0 license, see the license statement in the paper or at the digital library page of the publisher)
CC-BY 4.0 license, see the license statement in the paper or at the digital library page of the publisher)
Main Reference:
This work was done as a collaboration between the AVIZ project group of Inria, France, the MaiAGE team of InraE Jouy-en-Josas, and several other teams.
This work was supported through the Naviscope project, funded by Inria, France.