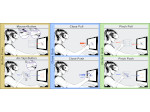
Multiscale Molecular Visualization
Description:
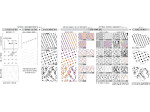
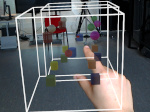
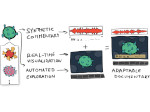
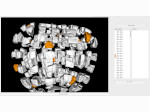
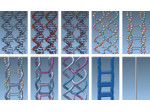
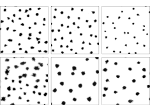
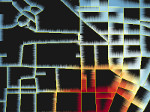
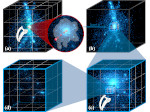
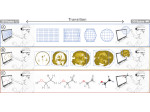
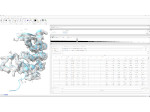
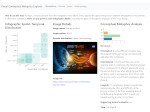
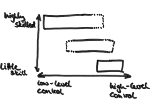
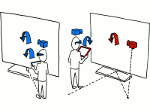
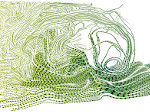
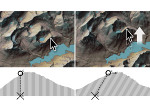
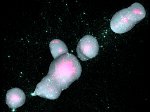
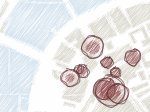
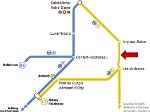
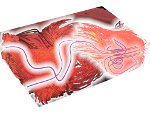
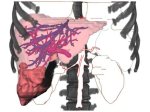
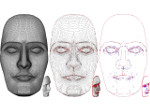
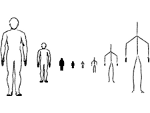
We provide a high-level survey of multiscale molecular visualization techniques, with a focus on application-domain questions, challenges, and tasks. We provide a general introduction to molecular visualization basics and describe a number of domain-specific tasks that drive this work. These tasks, in turn, serve as the general structure of the following survey. First we discuss methods that support the visual analysis of molecular dynamics simulations. We discuss, in particular, visual abstraction and temporal aggregation. In the second part we survey multiscale approaches that support the design, analysis, and manipulation of DNA nanostructures and related concepts for abstraction, scale transition, scale-dependent modeling, and navigation of the resulting abstraction spaces. In the third part of the survey we showcase approaches that support interactive exploration within large structural biology assemblies up to the size of bacterial cells. We describe fundamental rendering techniques as well as approaches for element instantiation, visibility management, visual guidance, camera control, and support of depth perception. We close the survey with a brief listing of important tools that implement many of the discussed approaches and a conclusion that provides some research challenges in the field.
Paper download:  (14.4 MB)
(14.4 MB)
Main Reference:
Other Reference:
| Haichao Miao (2019) Geometric Abstraction for Effective Visualization and Modeling. PhD thesis, TU Wien, Austria, August 2019. | | ||
This work was done in collaboration with the Research Unit of Computer Graphics at TU Wien, Austria, Masaryk University Brno, Czechia, the University of Stuttgart, Germany, and the VRVis Research Center, Austria.
This work was supported through the Illustrare project, funded by ANR in France and FWF in Austria.